Revision 18
#86652
Store at 4C and -20C
877-616-CELL (2355)
877-678-TECH (8324)
3 Trask Lane | Danvers | Massachusetts | 01923 | USA
For Research Use Only. Not for Use in Diagnostic Procedures.
Product Includes | Mini Kit (P) Quantity | Standard Kit (S) Quantity | Storage Temp |
|---|---|---|---|
Concanavalin A Magnetic Beads | 1 x 80 µl | 1 x 240 µl | +4°C |
Concanavalin A Bead Activation Buffer | 1 x 1.7 mL | 1 x 5 mL | +4°C |
1 x 800 µl | 1 x 2.5 ml | +4°C | |
1 x 4.5 mL | 1 x 15 ml | +4°C | |
1 x 3.6 mL | 1 x 7 ml | +4°C | |
Calcium Chloride | 1 x 25 µl | 1 x 100 µl | +4°C |
pAG-MNase Enzyme | 1 x 20 µl | 1 x 40 µl | -20°C |
1 x 1.2 mL | 2 x 1.2 ml | -20°C | |
1 x 300 µl | 1 x 1 ml | -20°C | |
1 x 400 µl | 1 x 1.3 ml | -20°C | |
1 x 200 µl | 1 x 750 µl | -20°C | |
1 x 20 µl | 1 x 100 µl | -20°C | |
1 x 20 µl | 1 x 50 µl | -20°C | |
1 x 20 µl | 1 x 20 µl | -20°C | |
1 x 20 µl | 1 x 100 µl | -20°C | |
Sample Normalization Spike-In DNA (1 ng/μl) | 1 x 40 µl | 1 x 120 µl | -20°C |
Sample Normalization Primer Set | 1 x 64 µl | 1 x 150 µl | -20°C |
1 x 50 µl | 1 x 150 µl | -20°C | |
1 x 50 µl | 1 x 150 µl | -20°C |
Applications:
C&R
Storage
Specificity/Sensitivity
The CUT&RUN Assay Kit can be utilized with any CUT&RUN-validated antibody to detect endogenous levels of protein-DNA interactions and histone modifications in mammalian cells (see Figures 1–6). The kit is compatible with multiple species of antibodies, including rabbit and mouse. The positive control Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751 detects multiple species of tri-methyl histone H3 Lys4 protein, including human, mouse, rat, and monkey. Primer sets are included for the human (#7014) and mouse (#7015) positive control RPL30 gene locus; however, the use of other species with the kit requires the design of additional control primer sets.
Description
The CUT&RUN Assay Kit is designed to conveniently provide reagents needed to perform up to 8 (P size) or 24 (S size) digestion reactions from cells and is optimized for 100,000 cells per reaction. The kit has been optimized to work for all types of DNA binding proteins, including histones, transcription factors and cofactors. A complete assay can be performed in as little as one day.
The CUT&RUN Assay Kit also provides important controls to ensure a successful CUT&RUN experiment. The kit contains a positive control Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751 and a negative control Rabbit (DA1E) mAb IgG XP® Isotype Control (CUT&RUN) #66362, both of which can be used for qPCR or Next Generation sequencing (NG-seq) analysis. PCR primer sets are provided for the human (#7014) and mouse (#7015) RPL30 gene locus to be used in conjunction with the control antibodies. This kit is compatible with both qPCR and NG-seq.
Background
Like the chromatin immunoprecipitation (ChIP) assay, Cleavage Under Targets & Release Using Nuclease (CUT&RUN) is a powerful and versatile technique used for probing protein-DNA interactions within the natural chromatin context of the cell (1-4). This assay can be used to identify multiple proteins associated with a specific region of the genome, or the opposite, to identify the many regions of the genome associated with a particular protein. In addition, the CUT&RUN assay can be used to define the spatial and temporal relationship of a particular protein-DNA interaction. For example, the CUT&RUN assay can be used to determine the specific order of recruitment of various protein factors to a gene promoter or to “measure” the relative amount of a particular histone modification across an entire gene locus during gene activation. In addition to histone proteins, the CUT&RUN assay can also be used to analyze binding of transcription factors and cofactors, DNA replication factors, and DNA repair proteins (Figures 1-6).CUT&RUN provides a rapid, robust, and true low cell number assay for detection of protein-DNA interactions in the cell. Unlike the ChIP assay, CUT&RUN is free from formaldehyde cross-linking, chromatin fragmentation, and immunoprecipitation, making it a much faster and more efficient method for enriching protein-DNA interactions and identifying target genes. CUT&RUN can be performed in less than one day, from live cells to purified DNA, and has been shown to work with as few as 500-1000 cells per assay (1,2). Instead of fragmenting all of the cellular chromatin as done in ChIP, CUT&RUN utilizes an antibody-targeted digestion of chromatin, resulting in much lower background signal than seen in the ChIP assay. As a result, CUT&RUN requires only 1/10th of the sequencing depth that is required for ChIP-seq assays (1,2). Finally, the inclusion of simple spike-in control DNA allows for accurate quantification and normalization of target-protein binding that is not possible with the ChIP method. This provides for effective normalization of signal between samples and between experiments.
Species Reactivity
Species reactivity is determined by testing in at least one approved application (e.g., western blot).
Applications Key
C&R: CUT&RUN
Cross-Reactivity Key
H: Human M: Mouse R: Rat Hm: Hamster Mk: Monkey Vir: Virus Mi: Mink C: Chicken Dm: D. melanogaster X: Xenopus Z: Zebrafish B: Bovine Dg: Dog Pg: Pig Sc: S. cerevisiae Ce: C. elegans Hr: Horse GP: Guinea Pig Rab: Rabbit G: Goat All: All Species Expected
Trademarks and Patents
Cell Signaling Technology is a trademark of Cell Signaling Technology, Inc.
SimpleChIP is a registered trademark of Cell Signaling Technology, Inc.
XP is a registered trademark of Cell Signaling Technology, Inc.
U.S. Patent No. 7,429,487.
All other trademarks are the property of their respective owners. Visit cellsignal.com/trademarks for more information.
Limited Uses
Except as otherwise expressly agreed in a writing signed by a legally authorized representative of CST, the following terms apply to Products provided by CST, its affiliates or its distributors. Any Customer's terms and conditions that are in addition to, or different from, those contained herein, unless separately accepted in writing by a legally authorized representative of CST, are rejected and are of no force or effect.
Products are labeled with For Research Use Only or a similar labeling statement and have not been approved, cleared, or licensed by the FDA or other regulatory foreign or domestic entity, for any purpose. Customer shall not use any Product for any diagnostic or therapeutic purpose, or otherwise in any manner that conflicts with its labeling statement. Products sold or licensed by CST are provided for Customer as the end-user and solely for research and development uses. Any use of Product for diagnostic, prophylactic or therapeutic purposes, or any purchase of Product for resale (alone or as a component) or other commercial purpose, requires a separate license from CST. Customer shall (a) not sell, license, loan, donate or otherwise transfer or make available any Product to any third party, whether alone or in combination with other materials, or use the Products to manufacture any commercial products, (b) not copy, modify, reverse engineer, decompile, disassemble or otherwise attempt to discover the underlying structure or technology of the Products, or use the Products for the purpose of developing any products or services that would compete with CST products or services, (c) not alter or remove from the Products any trademarks, trade names, logos, patent or copyright notices or markings, (d) use the Products solely in accordance with CST Product Terms of Sale and any applicable documentation, and (e) comply with any license, terms of service or similar agreement with respect to any third party products or services used by Customer in connection with the Products.
Revision 18
#86652
CUT&RUN Assay Kit
Figure 1. CUT&RUN was performed with 100,000, 10,000, or 5,000 HCT 116 cells (as indicated) and Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751, using the CUT&RUN Assay Kit. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. This figure shows binding of H3K4me3 across chromosome 12.
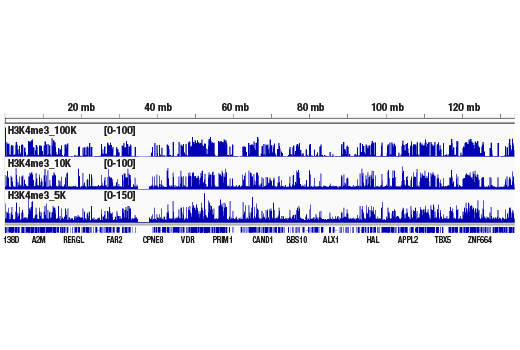
Figure 2. CUT&RUN was performed with 100,000, 10,000, or 5,000 HCT 116 cells (as indicated) and Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751, using the CUT&RUN Assay Kit. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. This figure shows enrichment around the GAPDH gene, a known target of H3K4me3.
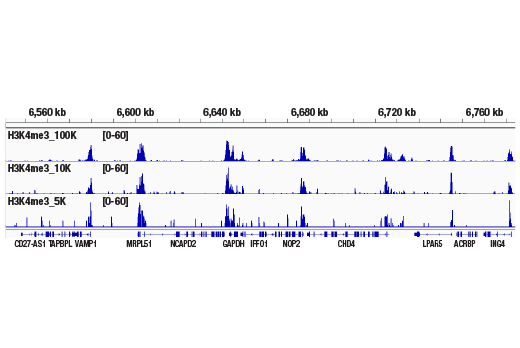
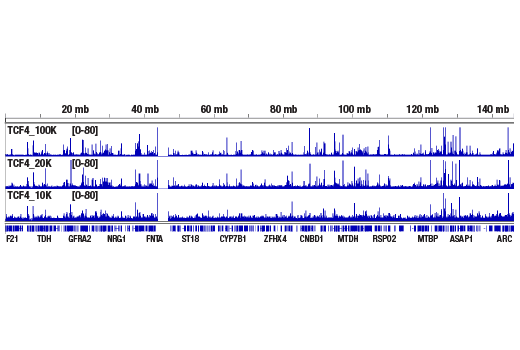
Figure 3. CUT&RUN was performed with 100,000, 20,000, or 10,000 HCT 116 cells (as indicated) and TCF4/TCF7L2 (C48H11) Rabbit mAb #2569, using the CUT&RUN Assay Kit. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. This figure shows binding of TCF4/TCFL2 across chromosome 8.
Revision 18
#86652
CUT&RUN Assay Kit
Figure 4. CUT&RUN was performed with 100,000, 20,000, or 10,000 NCCIT cells (as indicated) and SUZ12 (D39F6) XP® Rabbit mAb #3737, using the CUT&RUN Assay Kit. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. The figure shows binding across the HoxD gene cluster, a known target of SUZ12.
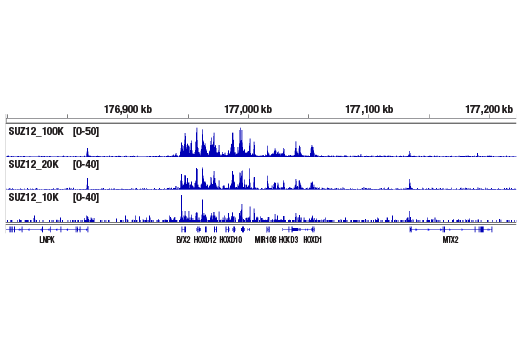
Figure 5. CUT&RUN was performed with 2.5 mg of medium fixed mouse liver tissue (0.1% formaldehyde, 10 min) and Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751, Glucocorticoid Receptor (D6H2L) XP® Rabbit mAb #12041, SUZ12 (D39F6) XP® Rabbit mAb #3737, or RING1B (D22F2) XP® Rabbit mAb #5694, using the CUT&RUN Assay Kit. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. The upper panel shows binding of H3K4me3 and Glucocorticoid Receptor around the Zfp866 gene, while the lower panel shows binding of SUZ12 and RING1B across the HoxA gene cluster.
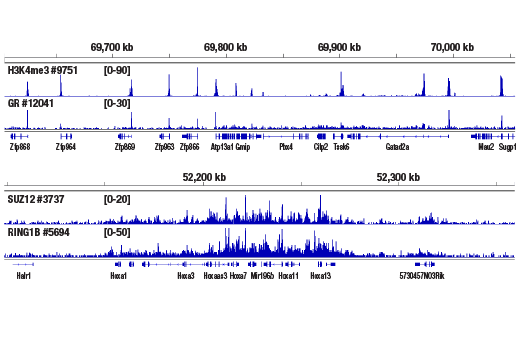
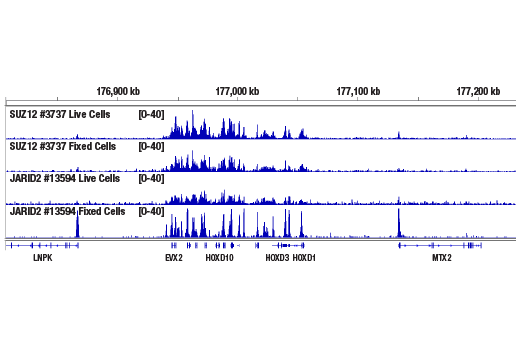
Figure 6. CUT&RUN was performed with 100,000 live or fixed NCCIT cells and SUZ12 (D39F6) XP® Rabbit mAb #3737 or JARID2 (D6M9X) Rabbit mAb #13594, using the CUT&RUN Assay Kit. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. The figure shows binding of SUZ12 and JARID2 across the HOXD gene cluster. Fixation is not required for SUZ12 (a core component of PRC2 complex) but increases the enrichment for JARID2 (an accessory component of PRC2 complex).
Revision 18
#86652
CUT&RUN Assay Kit
Figure 7. CUT&RUN and ChIP assays were performed with HCT 116 cells and Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751 or Rabbit (DA1E) mAb IgG XP® Isotype Control (CUT&RUN) #66362, using this CUT&RUN Assay Kit (left panel) or the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Magnetic Beads) #9005 (right panel). The enriched DNA was quantified by real-time PCR using SimpleChIP® Human GAPDH Exon 1 Primers #5516 and SimpleChIP® Human α Satellite Repeat Primers #4486. The amount of immunoprecipitated DNA in each sample is represented as signal relative to the total amount of input chromatin, which is equivalent to one.
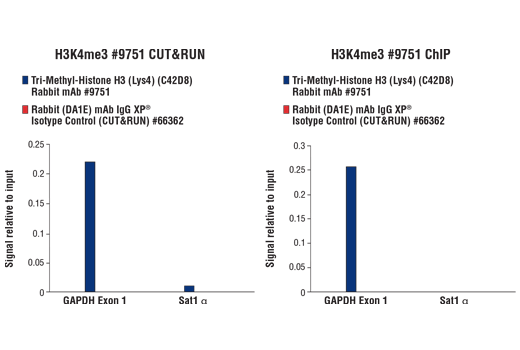
Figure 8. CUT&RUN and ChIP assays were performed with HCT 116 cells and Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751, using this CUT&RUN Assay Kit or the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Magnetic Beads) #9005. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. The upper panel compares enrichment of H3K4me3 across chromosome 12, while the lower panel compares enrichment at the GAPDH gene, a known target of H3K4me3. The input tracks are from the CUT&RUN input sample.
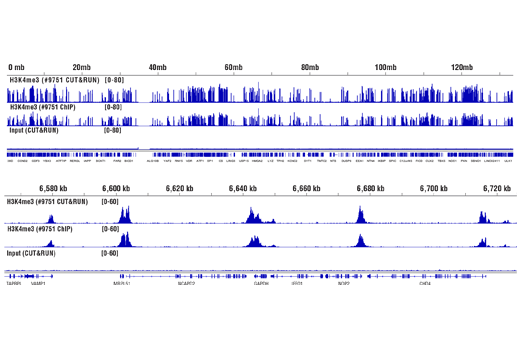
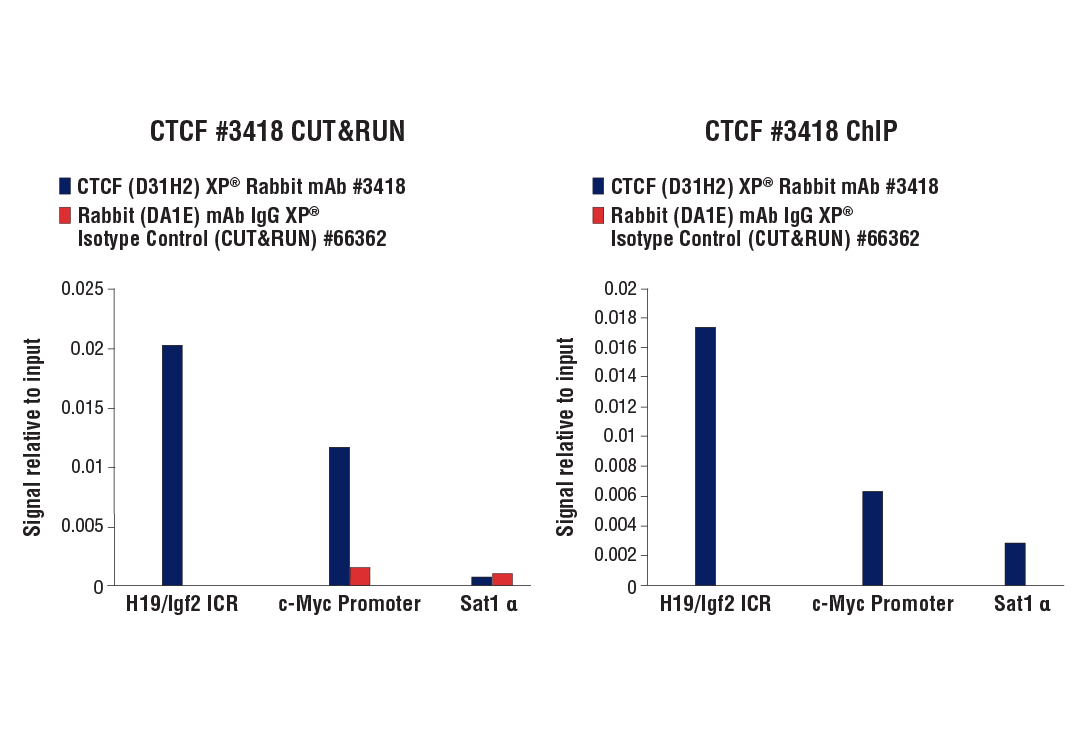
Figure 9. CUT&RUN and ChIP assays were performed with HCT 116 cells and CTCF (D31H2) XP® Rabbit mAb #3418 or Rabbit (DA1E) mAb IgG XP® Isotype Control (CUT&RUN) #66362, using this CUT&RUN Assay Kit (left panel) or the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Magnetic Beads) #9005 (right panel). The enriched DNA was quantified by real-time PCR using human c-Myc promoter primers, SimpleChIP® Human H19/Igf2 ICR Primers #5172, and SimpleChIP® Human α Satellite Repeat Primers #4486. The amount of immunoprecipitated DNA in each sample is represented as signal relative to the total amount of input chromatin, which is equivalent to one.
Revision 18
#86652
CUT&RUN Assay Kit
Figure 10. CUT&RUN and ChIP assays were performed with HCT 116 cells and CTCF (D31H2) XP® Rabbit mAb #3418, using this CUT&RUN Assay Kit or the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Magnetic Beads) #9005. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. The upper panel compares enrichment of CTCF across chromosome 8, while the lower panel compares enrichment at the MYC gene, a known target of CTCF. The input tracks are from the CUT&RUN input sample.
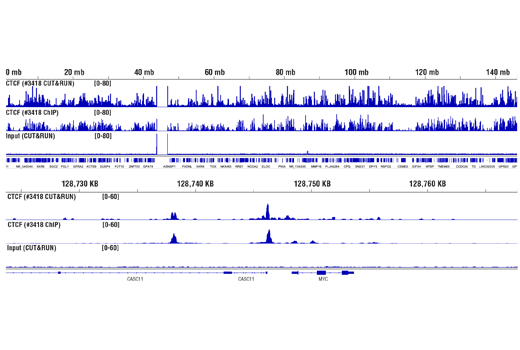
Figure 11. CUT&RUN and ChIP assays were performed with HeLa cells and Rpb1 CTD (4H8) Mouse mAb #2629 or Rabbit (DA1E) mAb IgG XP® Isotype Control (CUT&RUN) #66362, using this CUT&RUN Assay Kit (left panel) or the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Magnetic Beads) #9005 (right panel). The enriched DNA was quantified by real-time PCR using SimpleChIP® Human RPL30 Exon 3 Primers #7014, SimpleChIP® Human GAPDH Exon 1 Primers #5516, and SimpleChIP® Human MyoD1 Exon 1 Primers #4490. The amount of immunoprecipitated DNA in each sample is represented as signal relative to the total amount of input chromatin, which is equivalent to one.
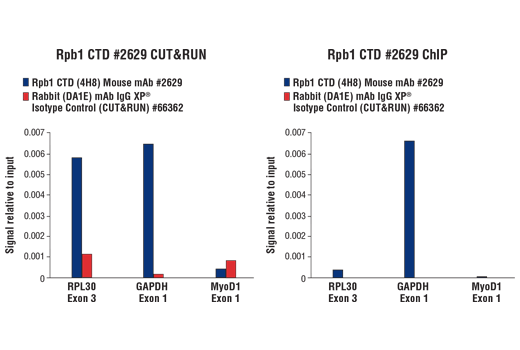
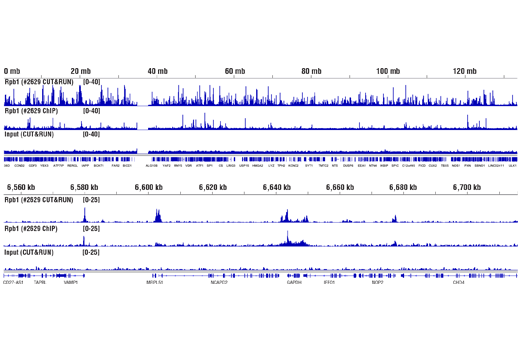
Figure 12. CUT&RUN and ChIP assays were performed with HeLa cells and Rpb1 CTD (4H8) Mouse mAb #2629, using this CUT&RUN Assay Kit or the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Magnetic Beads) #9005. DNA Libraries were prepared using DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795. The upper panel compares enrichment of Rpb1 across chromosome 12, while the lower panel compares enrichment at the GAPDH gene, a known target of Rbp1. The input tracks are from the CUT&RUN input sample.